|
Our group uses and develops Nuclear Magnetic Resonance (NMR) methods to study fundamental biophysical properties of proteins. While a wealth of data has been amassed over the course of years, many fundamental properties of proteins remain poorly understood. For instance, a protein structure cannot be reliably predicted based on a primary sequence. Relatively little is known about folding mechanisms. A vast class of loosely structured (unfolded) proteins has not been properly characterized. Considering protein-protein interactions, little information is available about short-lived transition states. Structural determinants of protein stability remain elusive. The list goes on.
|
 |
The main thrust of our work is to design NMR experiments and to develop associated theory in order to contribute to these important areas.
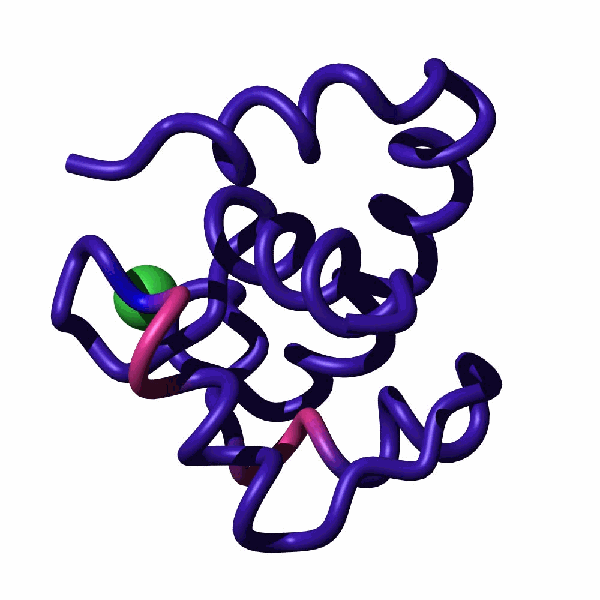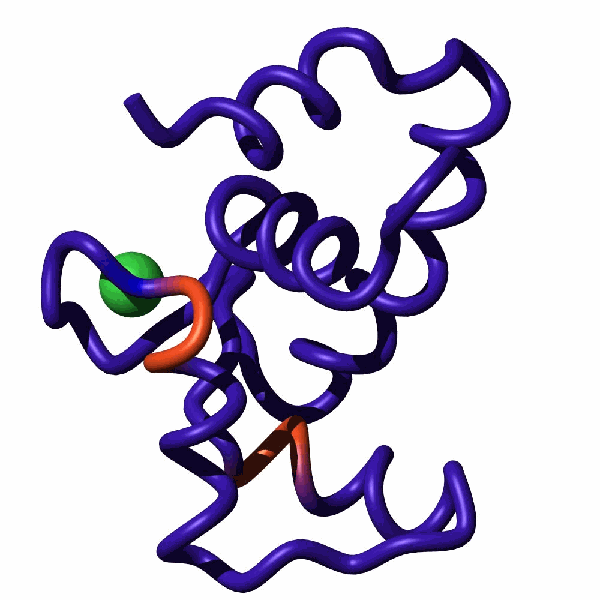
Cardiac troponin C (cTnC) is a part of the calcium-regulated signaling cascade responsible for heart beat. When loaded with a Ca2+ ion, the N-terminal domain of cTnC establishes dynamic equilibrium involving the closed and open forms. The open form is primed for binding a partner protein, troponin I, paving the way for a stream of events that ultimately leads to contraction of the heart muscle. The detection of a sparsely populated open form of NcTnC represents a considerable challenge. In our study, we demonstrated that replacing the Ca2+ ion with its mimetic, Ln3+, allows to highlight the dynamic exchange between the open and closed forms. Specifically, long-range pseudocontact shifts induced by Ln3+ ions enhance the amplitude of relaxation dispersion profiles reporting on µs-ms time-scale dynamics.
|
|
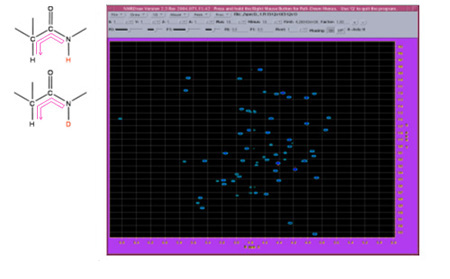
Backbone 15N relaxation is a principal source of information on protein dynamics. An estimated seven hundred research articles report on protein 15N relaxation, drawing a connection to protein function, structure, stability, binding affinity, etc. In our work we expanded the scope of nitrogen relaxation studies by targeting 15N-2H amide groups along with the standard 15N-1H. As it turns out, the data obtained from the deuterated sites are uniquely sensitive to the slower forms of internal dynamics that occur on a time scale of ~1 ns.
|
 |
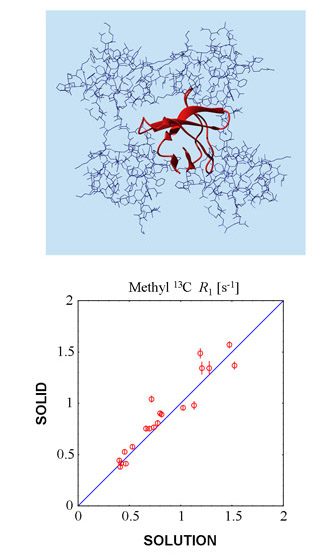 |
While solution-state NMR is a well-established tool for studies of large biomolecules, solid-state NMR is currently on the way to acquiring similar capabilities. We joined forces with the group of Prof. Bernd Reif in Berlin seeking to combine the data from these two techniques. Of note, protein samples used in solid- and solution-state NMR are fundamentally similar: in both cases the protein environment is mainly constituted by water, with occasional protein-protein contacts. Spectroscopically, however, the two situations are quite different. In solution, the relaxation data are sensitive only to faster forms of internal motion (since slower modes are screened out by the overall protein tumbling). In solids, both faster and slower forms of motion make their presence felt (proteins are held in place by the crystalline matrix and do not tumble). The combination of the two types of data - one selectively sensitive to picosecond motions and the other broadly sensitive to picosecond-nanosecond motions - provides a unique opportunity for comprehensive characterization of protein dynamics. We have used an array of spin probes, such as backbone 15N as well as side-chain 13C and 2H, to apply, develop, and test these concepts.
|
|
According to some estimates, 30% of proteins in eucaryotic proteome contain long stretches of disordered polypeptide chain. While seemingly random, many of these fragments possess an uncanny ability to recognize their unique binding partners. The structural basis for this specificity remains poorly understood. Until now disordered proteins remain a grey area on the otherwise fairly well charted proteome map.
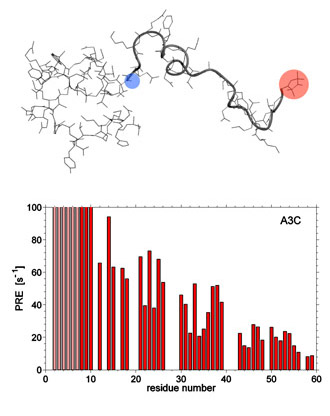
Among many spectroscopic techniques used to probe unfolded proteins, NMR paramagnetic relaxation enhancement (PRE) experiment is arguably the most promising. Surprisingly, the proper theory of the PRE effect is unfolded protein has so far been lacking. To address this problem, we developed a number of models for relative motion of paramagnetic label and the reporter 1HN spins, including analytical, numeric, and MD based models. We have also improved the experimental procedure by introducing the reference compound and by focusing on peak volumes instead of peak intensities. The application to denatured drkN SH3 domain showed the preponderance of extended conformations, yet also produced the evidence of compact (collapsed) species.
|
 |
|
A number of other projects are underway in the lab: we are conducting molecular dynamics simulations of proteins constrained by crystal lattice, refine a protein structure solved by solid-state NMR, investigate dynamics of protein domains connected via long flexible linker, explore protein solvation and internal motions in water-glycerol solvent, quantitate amide exchange in unfolded proteins using a newly developed pulse sequence, test new paramagnetic tags for studies of unfolded proteins, etc.
|
|

