Andrew Mesecar

- Distinguished Walther Professor of Cancer Structural Biology | Biochemistry
- Email: amesecar@purdue.edu
- Phone: 41924
- Office: 311 HOCK
The main interest of the Mesecar lab is the Structure & Function of Enzymes of biomedical importance. One of our fundamental goals is to gain a deeper understanding of the roles these enzymes play in the disease process and how these enzymes recognize and catalyze the transformation of their substrates into products. By understanding enzymes’ molecular recognition processes and their kinetic, chemical, structural and regulatory mechanisms, we can then exploit this knowledge for the discovery, design and development of small molecule drugs. We are currently studying the structure and function of enzymes involved in promoting cancer, coronavirus infections and Alzheimer’s disease.
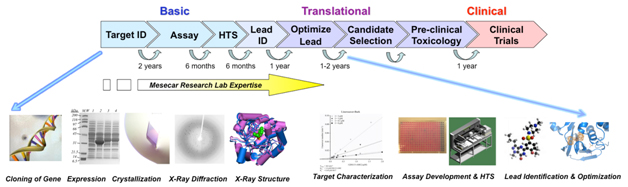
Due to the complex nature of enzymatic catalysis and molecular recognition, we routinely use a variety of state-of-the-art experimental approaches and tools from the fields of chemistry, biology and physics. Our main experimental tools are static and time-resolved X-ray crystallography, enzyme chemistry & kinetics, molecular biology, mass-spec proteomics, assay development and optimization, high-throughput screening and molecular modeling, cheminformatics and cryo-EM. Our multi-faceted, multi-disciplinary, collaborative and integrated approach to the elucidation of the nature of enzymatic catalysis and molecular recognition is the model for scientific investigation and education in the twenty-first century.
Education
- B.S., Purdue University, Chemistry (ACS), 1988
- Ph.D., University of Notre Dame, Biochemistry, 1995
- Post-Doctorate, University of California, Berkeley, 1998
Awards
- Outstanding Graduate Educator Award - Purdue University Office of Provost, 2021
- Defense Science Study Group - U.S. Department of Defense, 2006
- American Association of Colleges of Pharmacy New Investigator Award, 1999
- U.S. Department of Energy: Post-Doctoral Fellowship Center for Advanced Materials, Lawrence Berkeley National Laboratory, 1995
- Reilly Graduate Fellowship: University of Notre Dame, 1993
- U.S. Department of Education: Graduate Fellowship. GAANNP Scholarship Program, 1989
- U.S. Air Force ROTC Four-Year Scholarship, 1984